| 別品名 |
Core-binding factor subunit beta, CBF-beta, Polyomavirus enhancer-binding protein 2 beta subunit, PEA2-beta, PEBP2-beta, SL3-3 enhancer factor 1 subunit beta, SL3/AKV core-binding factor beta subunit
|
| 種由来 |
Human
|
| 標識物 |
Unlabeled
|
| 精製度 |
Serum
|
| 適用 |
Enzyme Linked Immunosorbent Assay
Chromatin Immunoprecipitation
|
| 免疫動物 |
Rabbit
|
| 抗体クラス |
IgG1
|
| 交差種 |
Human
|
| GENE ID |
865
|
| Accession No.(Gene/Protein) |
NP_001746, Q13951
|
| Gene Symbol |
CBFB
|
| 形状 |
滅菌済み液状品
|
| 参考文献 |
Martens JHA, Mandoli A, Simmer F, Wierenga B-J, Saeed S, Singh AA, Altucci L, Vellenga E, Stunnenberg HG (2012) ERG and FLI1 binding sites demarcate targets for aberrant epigenetic regulation by AML1-ETO in acute myeloid leukemia. Blood 120: 4038-4048.
|
|
※サムネイル画像をクリックすると拡大画像が表示されます。
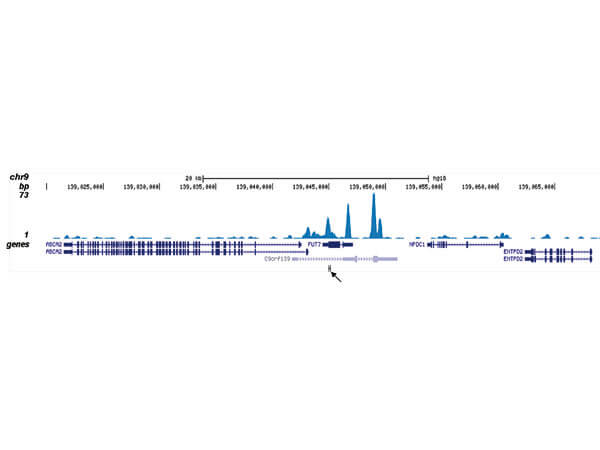
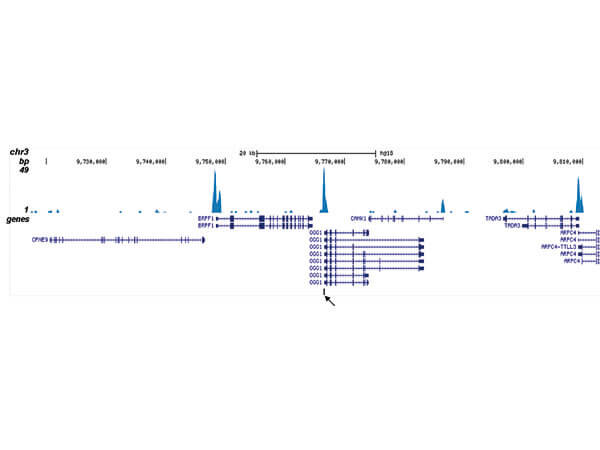
ChIP seq results obtained with the antibody directed against CBFb. ChIP was performed as described in figure 1. The IPd DNA from 6 ChIPs were pooled and analysed with an Illumina Genome Analyzer. Library preparation, cluster generation, and sequencing were performed according to the manufacturers instructions. The 32 bp tags were aligned to the human reference genome (hg18) using the ELAND algorithm. Figure 2 shows the results of the complete chromosome 3. Figure 3 5 shows three genomic regions region surrounding the OGG1, FUT7 and NFE2 genes, respectively. The position of the PCR amplicon is indicated with an arrow.
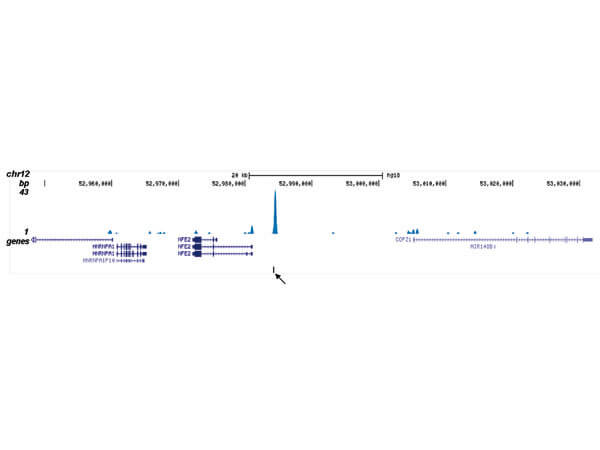
ChIP-seq results obtained with the antibody directed against CBFb. ChIP was performed as described in figure 1. The IPd DNA from 6 ChIPs were pooled and analyzed with an Illumina Genome Analyzer. Library preparation, cluster generation, and sequencing were performed according to the manufacturers instructions. The 32 bp tags were aligned to the human reference genome (hg18) using the ELAND algorithm. Figure 2 shows the results of the complete chromosome 3. Figure 3-5 shows three genomic regions region surrounding the OGG1, FUT7 and NFE2 genes, respectively. The position of the PCR amplicon is indicated with an arrow.
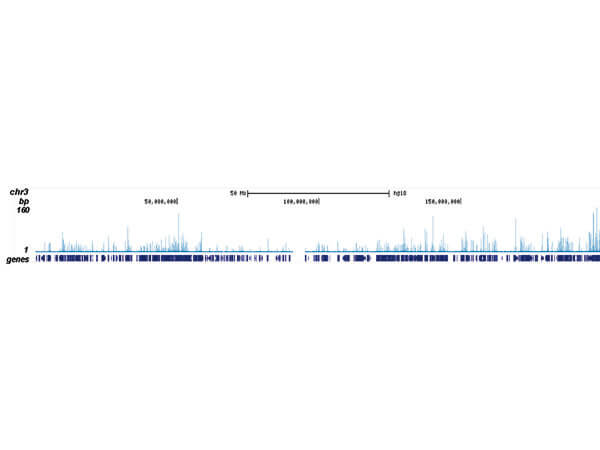
ChIP-seq results obtained with the antibody directed against CBFb. ChIP was performed as described in figure 1. The IPd DNA from 6 ChIPs were pooled and analysed with an Illumina Genome Analyzer. Library preparation, cluster generation, and sequencing were performed according to the manufacturers instructions. The 32 bp tags were aligned to the human reference genome (hg18) using the ELAND algorithm. Figure 2 shows the results of the complete chromosome 3. Figure 3-5 shows three genomic regions region surrounding the OGG1, FUT7 and NFE2 genes, respectively. The position of the PCR amplicon is indicated with an arrow.
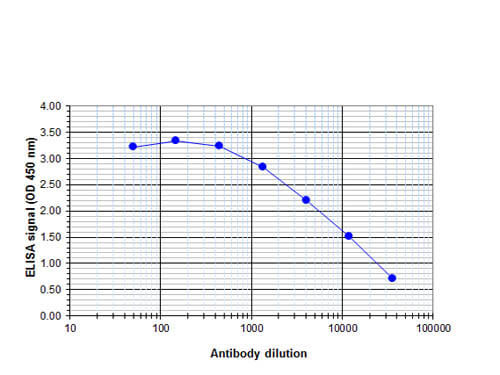
ELISA results of Rabbit anti-human CBFb antibody. Antigen: BSA conjugated CBFb. Coating amount: 0.1 ug per well. Dilution series: serial dilution. Estimated Antibody Titer to be 1:8,800. Substrate: TMB (p/n TMBE-1000).
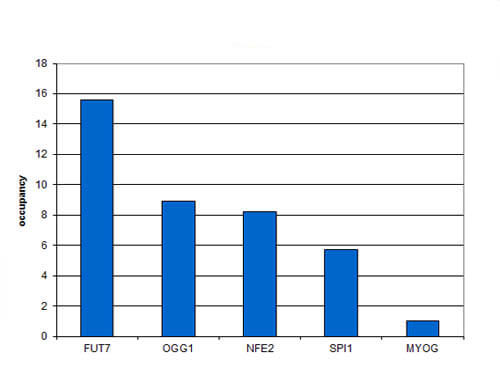
Chromatin Immunoprecipitation results of Rabbit Anti-CBFb Antibody. Chromatin from 1.25 million formaldehyde cross-linked SKNO-1 cells was used with 4ul of Anti-human CBFb Antibody and 20ul of magnetic IgG beads per immunoprecipitation. QPCR was performed using primers specific for the FUT7, OGG1, NFE2, and SPI1 genes. ChIP results shows the relative occupancy, calculated as the ratio + control/background for which the MYOG gene was used.
ChIP-seq results obtained with the antibody directed against CBFb. ChIP was performed as described in figure 1. The IPd DNA from 6 ChIPs were pooled and analysed with an Illumina Genome Analyzer. Library preparation, cluster generation, and sequencing were performed according to the manufacturers instructions. The 32 bp tags were aligned to the human reference genome (hg18) using the ELAND algorithm. Figure 2 shows the results of the complete chromosome 3. Figure 3-5 shows three genomic regions region surrounding the OGG1, FUT7 and NFE2 genes, respectively. The position of the PCR amplicon is indicated with an arrow.
|

|
|
ChIP seq results obtained with the antibody directed against CBFb. ChIP was performed as described in figure 1. The IPd DNA from 6 ChIPs were pooled and analysed with an Illumina Genome Analyzer. Library preparation, cluster generation, and sequencing were performed according to the manufacturers instructions. The 32 bp tags were aligned to the human reference genome (hg18) using the ELAND algorithm. Figure 2 shows the results of the complete chromosome 3. Figure 3 5 shows three genomic regions region surrounding the OGG1, FUT7 and NFE2 genes, respectively. The position of the PCR amplicon is indicated with an arrow.
|
|
|
| メーカー |
品番 |
包装 |
|
RKL
|
100-401-V32
|
100 UL
|
※表示価格について
| 当社在庫 |
なし
|
| 納期目安 |
約10日
|
| 保存温度 |
-20℃
|
|